
 Jessica passed her oral exams this morning! If you weren't able to make it to the celebration, be sure to congratulate her next time you see her. We had a champagne toast in the lab, followed by lunch at Phil's Sliders
0 Comments
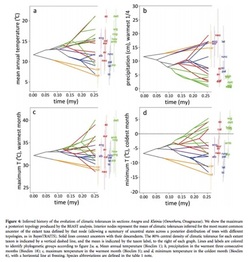 The other day I got into a class discussion about ancestral state reconstruction, or the assignment of ancestral states based on extant taxa. Some people thought that character mapping was not scientifically sound unless you had a time machine. I, however, disagree. The utility of phylogenetics is to determine evolutionary relationships between extant taxa, and while time machines do not exist, systematists are able to make hypotheses about these relationships. I think character mapping follows the same principles and gives science more insight into diversification processes. However, one problem I do see with ancestral state reconstruction is that your analysis is only as good as your phylogenetic tree. As many people say, “junk in, junk out.” Therefore, use with caution. One new ancestral state reconstruction method I recently learned uses the software package Phyloclim maintained by Christoph Heibl. Phyloclim is a package implemented in R that integrates species ecological niche models with phylogenetics in order to calculate and visualize niche evolution on a phylogenetic tree. To do this, the user must first create a predicted species occupancy model. This involves using ecological niche modeling software, such as Maxent, and species location data stored as a raster file--I believe this is done in a mapping program like GIS. There is a great tutorial on the Maxent website. Patterns of niche evolution or niche conservation can then be determined by comparing the variation between niche space within and between subclades. One of two outcomes is expected: 1) either niche evolution occurs between subclades with conservation of niche space within each subclade or 2) niche conservation occurs between subclades and niche evolution occurs within each subclade. This means you could either see a divergence of niche spaces at the beginning family followed by conservation of their niche spaces or you would see niche space preserved early on in evolutionary history followed by diversification among species. Inference can then be made about ancestral niche spaces using a statistical analysis, and a statement of how climate (or other variables) may or may not result in the diversification of a group. I really liked using this R package. The visual of ancestral state reconstruction (see figure above) is really informative and neat to look at. However, Phyloclim is still quite new and no formal tutorial from the developers exists, but there are examples in the software package complete with example data. I have yet to try using my own data in Phyloclim, but I think it’s as simple reading a .csv file of your ecological niches models and your tree file in newick or nexus format (I like this tutorial) into R. I realize this may be easier said then done, but in theory it’s simple. Jessica Craft Useful papers: Evans, M. E. K., S. A. Smith, R. S. Flynn, and M. J. Donoghue. 2009. Climate, niche evolution, anddiversification of the ’bird-cage evening primroses’ (Oenothera, sections Anogra and Kleinia). Am. Nat. 173: 225-240. Fitzpatrick, B.M & Turelli, M. 2006. The geography of mammalian speciation:mixed signals from phylogenies and range maps. Evolution 60: 601-615. Phillips, S.J, M. Dudik, & R.E. Schapire. 2006. Maximum entropy modeling of species geographic distributions. Ecological Modeling 190: 231-259. Warren, D., R.E. Glor, & M. Turelli. 2008. Environmental niche equivalency versus conservatism: quantitative approaches to niche evolution. Evolution 62: 2868-2883 As a fly enthusiast, I understand how daunting a task identifying species can be. The minute details, the crazy terms: it can all make you lose your head, especially when you’ve gathered a seemingly infinite amount of specimens. But, what’s a scientist to do?
You could hunker down at a microscope and wait until your eyes cross, or you could head down the road of genetic barcoding. Now, simmer down, you taxonomists. I don’t plan to argue you guys out of your jobs. In fact, I have my own criticisms of barcoding, but just humor me for a moment. Genetic barcoding works by sequencing small DNA portions from unknown organisms and comparing those sequences to a barcode library. So say you’ve collected a bunch of something, let’s say unicorns from the North Pole as everyone knows all magical ponies live in the wintery north. Well, as a well-known unicorn scientist you are aware that there are several cryptic species of unicorns. This means that two or more species appear morphologically similar but, by at least one of the many species concepts, are still considered separate species. A quick PCR analysis, PCR gods forgiving, and a BLAST to the NCBI database could tell you which mythical unicorn species you now possess (should the barcode library of unicorns be complete). Okay, I may have lied. Unicorns don’t really exist (outside the imagination of yours truly), but the problem of cryptic species does, along with a myriad of other identification issues such as morphological variation within species and even between adults and juveniles. Have you ever looked at drosophila larvae? They all look like squiggly, little, wormy things, every single one of them. Aside from some neat distinguishing behaviors – a few fling themselves like trapeze artists – you couldn’t tell them apart. So, it makes sense that a useful tool like barcoding has received so much attention, but let’s not get carried away. This isn’t the messiah come here to solve all our problems. The way I see it genetic barcoding is the microwave of the 1970’s housewife: a new tool for the modern taxonomist. It heats your food in mere minutes, but you can still burn the pot roast. Criticisms include incorrectly identified species sequences, a substantial error rate, and lowered ability to distinguish between recently diverged species. These comments all point towards the necessity of well-studied taxonomists to make final decisions. Me? I’m sticking to the microscope for now. Having a good grasp on taxonomic identification seems like it will always be a useful tool. Jessica Craft Check out Jessica's paper reviewing a new book on Phylogenetic Networks in the journal Taxon. Congratulations!
While some insects can survive more than 10x the amount of radiation than humans, thank you MythBusters, the real inheritors of the nuclear apocalyptic world would have to be fungi. Why? Because radiation exposure for insects would continue after their initial exposure by seeping in through their food sources, but for a fungus that radiation would be lunch!
But how is this possible? First, the numbers of fungi inhabiting planet earth is immense. They are everywhere and you’ve probably experienced them in the form of dandruff or the dreaded athlete’s foot. So when black fungus was found growing on the walls of the still highly radioactive Chernobyl reactor, it wasn’t too exciting, until Dr. Auturo Casadevall began to wonder if the fungus was actually using the radiation as an energy/food source. “But I thought fungi were decomposers”, you might be saying to yourself, and you’d be right. Because fungi lack chlorophyll, fungi cannot synthesize their own energy source and therefore must consume organic compounds for fuel. Then how might fungus incorporate radiation, something very akin to sunlight, into their metabolism? The answer is melanin. Yup, I do mean the melanin in our skin that protects us from sunburns (or lack there of for our fair-skinned friends). However, fungi seem to use their melanin for energy capture rather than the acquisition of the perfect summer tan. Studies show that when the melanized fungal cells of Wangiella dermatitidis and Cryptococcus neoformans encounter 500 times the amount of background radiation they actually grow faster even under nutrition-limited conditions unlike their unmelaninized counterparts. So, when and if a nuclear holocaust does occur, the cockroaches will have plenty of food to munch on. In the meantime, science could utilize these melanin-rich fungi to expand our knowledge of energy capture in the human body, or even radiation clean-up, currently an unexplored arena. While I don’t expect fungi to sprout legs, don a spidey suit, and fight nuclear-aftermath crimes, I do think they are a start to recolonizing highly radioactive areas and that seems close enough to a super hero power. Jessica Craft First year graduate student Jessica Craft received honorable mention for her NSF GRFP and Ford Foundation proposals.
|
PatrickProfessor Archives
April 2018
Categories
All
|

 RSS Feed
RSS Feed
